
Our services
Molecular pathology platform
Our services
Molecular pathology platform
Genomic alterations of solid tumors from cancer patients can be detected in the laboratory by different techniques: Next-Generation Sequencing (Thermofischer PGM or Qiagen Genereader), Digital PCR (Stilla), Real-Time PCR (Idylla Biocartis, Cobas Roche or rotorgene Qiagen) Pyrosequencing (Q24 pyromark Qiagen), Direct Sequencing (ABI 3100 Applied) and FISH (Fluorescent In Situ Hybridization)
These techniques can be used both for personalized treatment of patients and for research projects.

Genomic profiling
Search for mutations from cells, tissues (frozen or embedded in paraffin) or directly from genomic DNA:
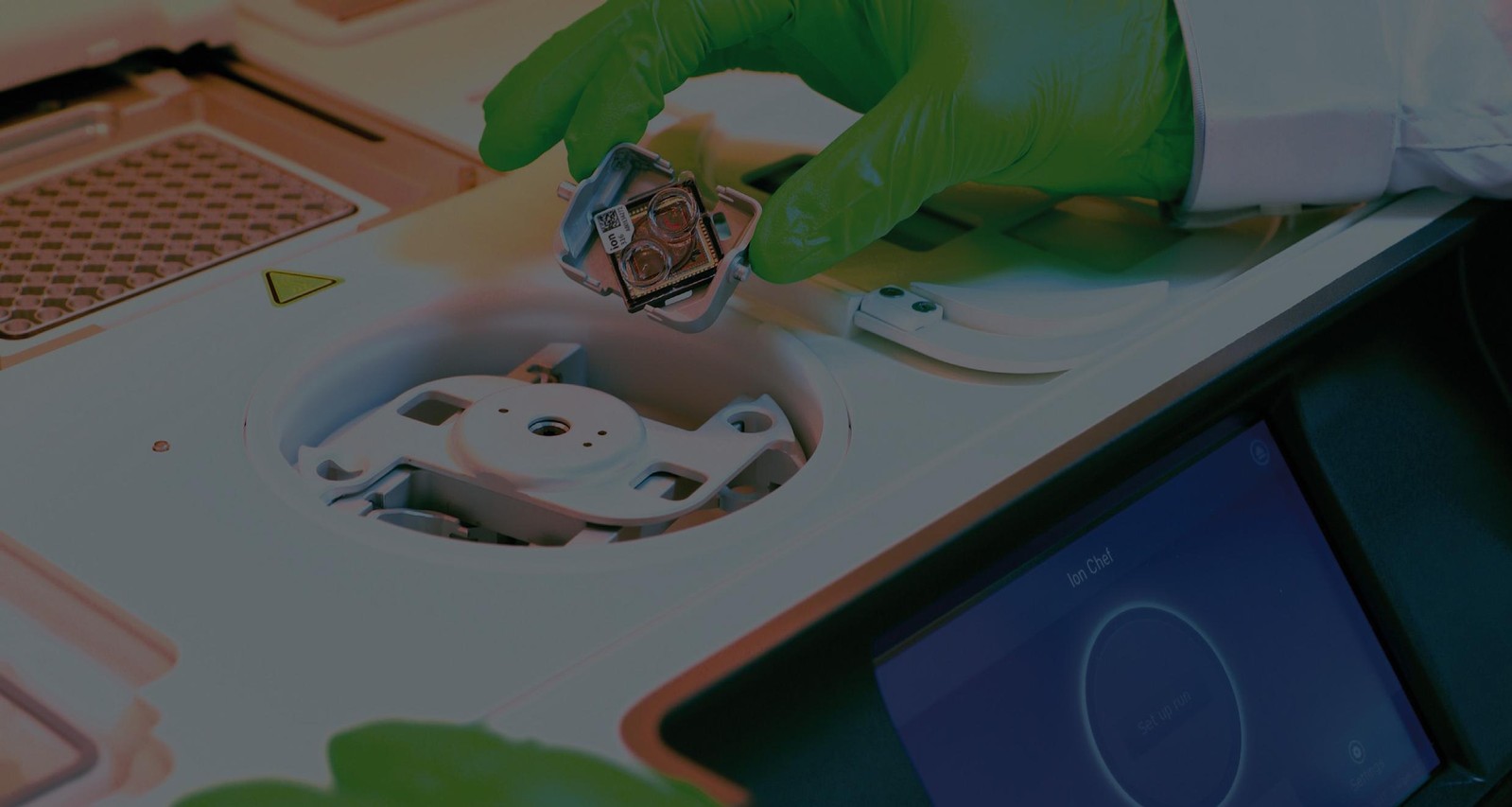
The Idylla technique (Biocartis) is utilized in our laboratory for several genes. It is a revolutionary approach that combines both DNA extraction and real-time PCR for a given gene. A whole range of genes have been commercialized by the company Biocartis and can be used in our laboratory.
PGM (Thermofisher) is used to look for somatic mutations in a broader spectrum by targeting 50 genes in cancer patients. This solution requires the users to conduct a bioinformatic interpretation of the data acquired during the sequencing.
The Genereader platform (QIAgen) includes various instruments, covering all stages, from DNA extraction to data analysis and interpretation, which can be edited as a report. This platform is used within the laboratory for cancer patients. The application targets somatic mutations on a panel of 12 genes (KRAS, NRAS, KIT, BRAF, PDGFRA, ALK, EGFR, ERBB2, PIK3CA, ERBB3, ESR1, RAF1). Other applications may be possible.
Pyrosequencing (Qiagen Technology) is used in our laboratory for targeted mutation research (EGFr, KRAS, BRAF, NRAS and any other mutation marketed by Qiagen; https://www.qiagen.com/fr/shop/detection-solutions/personalized-healthcare/). This is a fast technique that can sequence codons of interest from different genes. DNA extractions are performed with a kit (QIAamp DNA FFPE tissue for paraffin embedded tissue and MagNA Pure Compact Nucleic Acid Insulation for frozen tissues or for cells). The DNA is first amplified over the region of interest and then a pyrosequencing step targets the desired codons. Pyrograms are then obtained and analyzed by engineers of our laboratory, and authorized and validated by our molecular pathologists.
Sequencing by the Sanger method is performed in the laboratory with the 3500 Sequencer (Applied Biosystems). This technique makes it possible to sequence relatively long fragments. Once the DNA is extracted, it is amplified using specific primers that flank the desired area and the resulting PCR products are directly analyzed by the sequencer. The sequences thus obtained are examined by engineers from our laboratory and validated by our molecular pathologists.

Mutation status in liquid biopsy
Tumoral circulating DNA, present in the plasma of patients, is a valuable tool in thoracic oncology for the detection of Epidermal Growth Factor Receptor (EGFR) mutations. A simple blood sample (also called a liquid biopsy) can lead to a molecular profile diagnosis and also the early detection of disease relapse. A targeted therapy that exploits EGFR tyrosine kinase inhibitors is then proposed to the patient, as part of a personalized treatment.
At CRB LPCE, two techniques are used for the detection of EGFR mutations on plasmatic DNA: real-time PCR (Cobas® assay from Roche) and droplet-digital PCR (Naica® system by stilla). Cobas® assay (Roche) is based on plasmatic DNA extraction, followed by real-time PCR for amplification and detection of the mutations in exons 18, 19, and 20 of EGFR. The Naica® system (Stilla) focuses on digital PCR, a very sensitive and accurate technique for mutation detection and quantification, especially suitable for the analysis of tumoral DNA present in plasma at very low amounts. Nucleic acids are first fractionated in droplets, then separately amplified and analyzed. This very powerful technique takes into account each molecule of plasmatic DNA.
Gene rearrangements
Fluorescent In Situ Hybridization (FISH) is a molecular biology technique for the in situ detection of a target DNA sequence by hybridization with a complementary DNA probe linked to a fluorochrome.
In the laboratory, this technique makes it possible to demonstrate the rearrangement of the ALK gene. Any other gene rearrangement can also be considered.
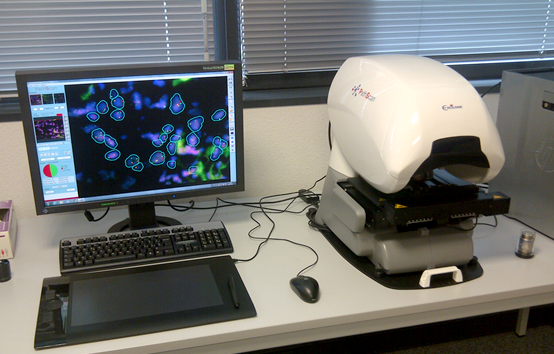
Hybridization is observed either by manual reading using a Nikon epifluorescence microscope or by the automated Pathscan Excilon platform for virtual slide FISH analysis by authorized personnel.