BIOBANKING
Dermatological
Collection
About
Dermatological collection
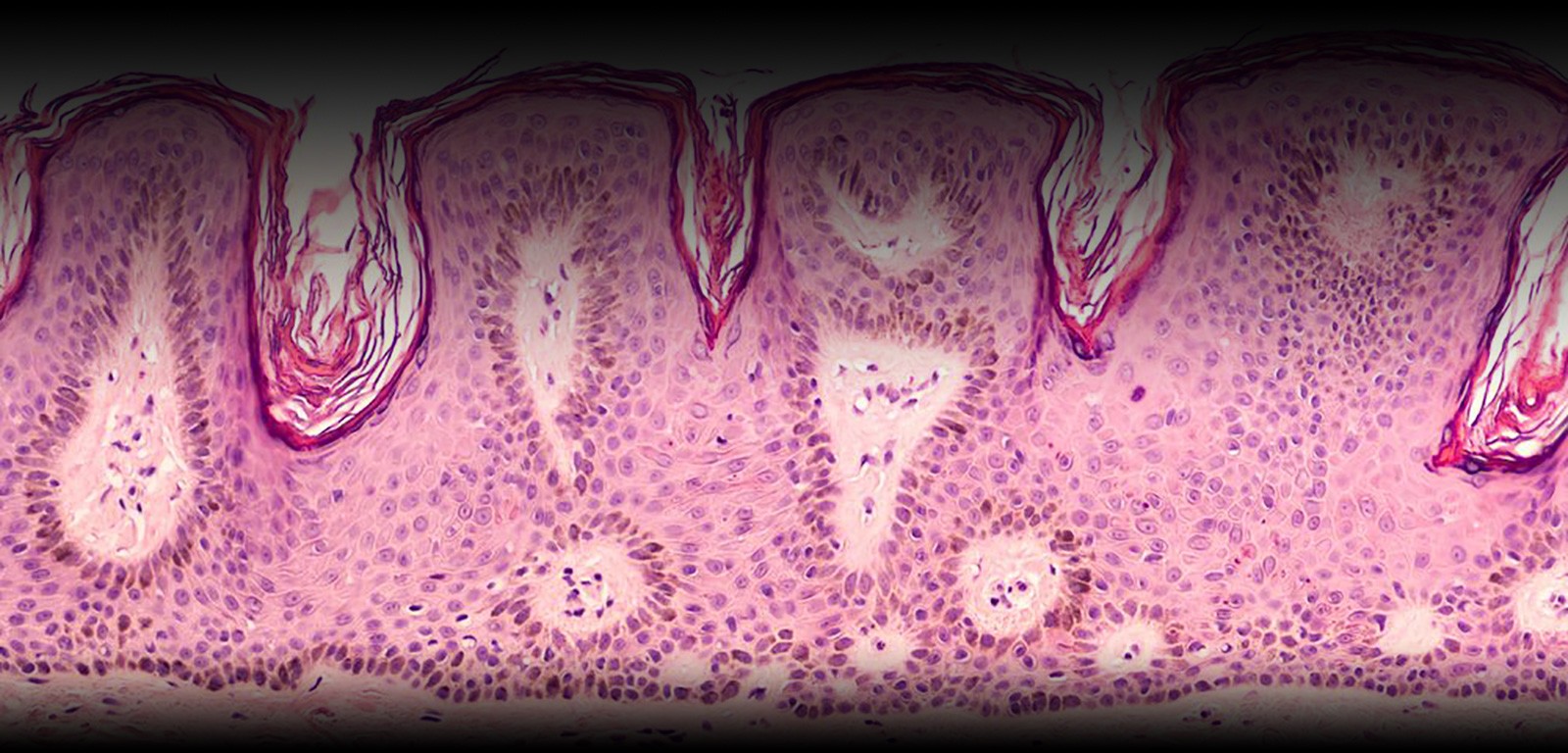
The dermatological collection essentially consists of melanoma specimens. Most of the samples originate from mucocutaneous melanoma with some ocular melanomas. As of 2018, it includes: 936 tissue samples and 674 blood samples, corresponding to 258 patients. These samples are associated with clinical data, including the identification of the primary site and the Breslow and Clark indexes.
Biosample diversity
For each patient, up to 4 fragments are collected from the lesion area and the healthy zone and stored at -80°C. For each tissue type, one of the fragments is dedicated to a diagnostic use, the others to research use.
Additional tissue fragments may be included in paraffin (room temperature storage) and some may be included in the tissue block collection (storage at 4°C). The paraffin-embedded tissue blocks can, on request, enable the construction of collections dedicated to targeted pathologies as tissue microarrays (TMA).
Characterization by immunohistochemistry of paraffin-embedded tissue blocks can be performed on request, with a whole range of antibodies.
Plasma and serum are collected from patient blood samples and stored at -80°C. Derivatives are also prepared from the samples: DNA and RNA from healthy and lesional tissues.
Identification of genetic alterations from tissues, blood samples or directly from nucleic acids can be performed on request.
Our
Services
Sample Transfer
The Biobank Côte d’Azur offers a full range of bioservices to industry, academia and international consortia.
Our policy is to be able to respond to different types of scientific and medical collaborations depending on the needs of applicants and the collaborative and cont[...]
Platforms
The Biobank Côte d’Azur collects human samples associated with various organs and pathologies.
For each of these, we collect numerous biosamples such as tissue fragments (lesional and peri-lesional tissues), biofluids and derived bioproducts. Each sample is associa[...]